The research, led by Michael Wagner from DOME in collaboration with CeMESS members and international partners from the UK, Denmark, and the US, highlights the often-overlooked effects of human-targeted drugs on microbial communities. Using a novel experimental approach, the team discovered that entacapone triggers iron starvation in the gut, promoting the growth of bacteria adapted to low-iron conditions. This groundbreaking study offers insights into how certain drugs can unintentionally alter the gut microbiome and suggests strategies to mitigate these effects.
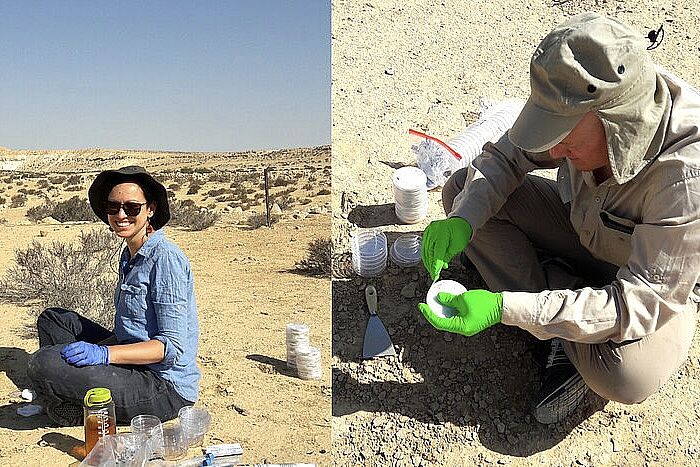
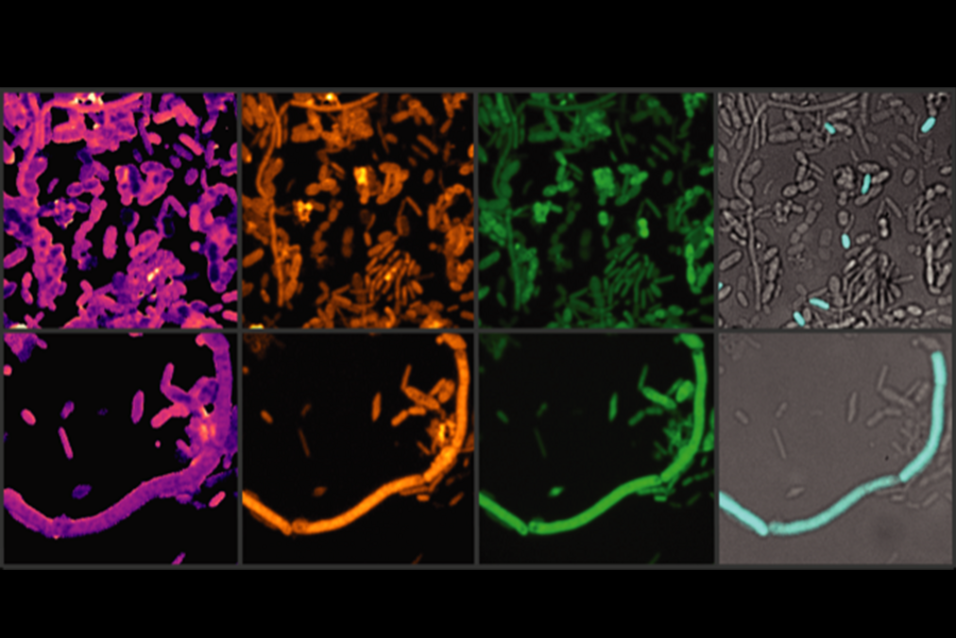
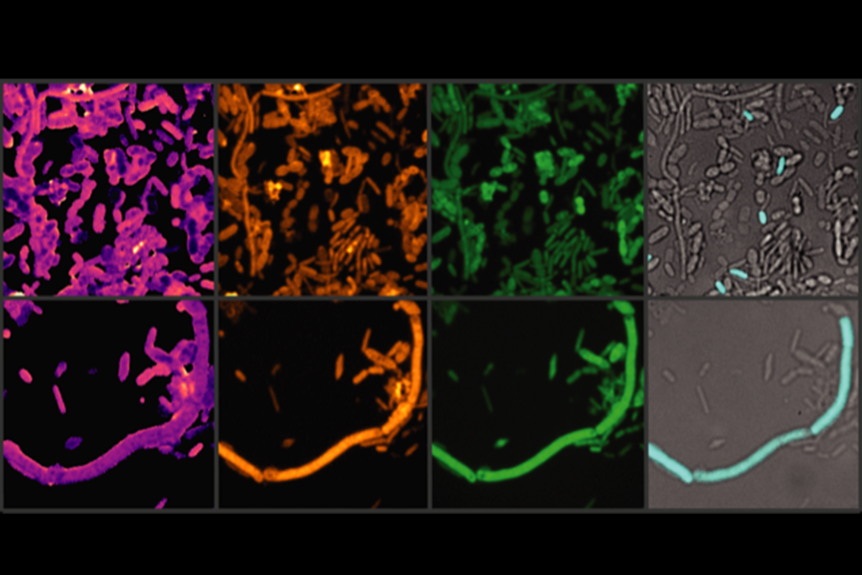
NanoSIMS was used in the study to analyze the iron content in gut microbiota cells after exposure to entacapone. The method involved preparing faecal and bacterial cell samples, which were fixed, washed, and filtered onto gold-coated polycarbonate filters. These were then analyzed using a NanoSIMS 50L ion microprobe. The technique enabled the detection of secondary ion signals for elements such as carbon (12C+), sodium (23Na+), phosphorus (31P+), calcium (40Ca+), and iron (56Fe+). The study used NanoSIMS imaging to generate high-resolution elemental distribution maps, revealing that entacapone exposure increased iron accumulation in gut bacteria, particularly in Phocaeicola dorei. The relative iron content was quantified by normalizing the 56Fe+ signal to 12C+, allowing single-cell level comparisons.